2.4.2.30: NAD+ ADP-ribosyltransferase
This is an abbreviated version!
For detailed information about NAD+ ADP-ribosyltransferase, go to the full flat file.
Reaction
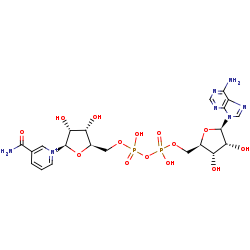 NAD+ +
NAD+ +
 (ADP-D-ribosyl)n-acceptor=
(ADP-D-ribosyl)n-acceptor=
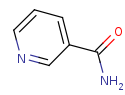 nicotinamide +
nicotinamide +
 (ADP-D-ribosyl)n+1-acceptor +
(ADP-D-ribosyl)n+1-acceptor +
 H+
H+
Synonyms
(adenosine diphosphoribose)transferase, nicotinamide adenine dinucleotide-protein, 193-kDa vault protein , 2,3,7,8-tetrachlorodibenzo-p-dioxin poly(ADP-ribose) polymerase, adenosine diphosphate ribosyltransferase, ADP-ribosylating thermozyme, ADP-ribosyltransferase, ADP-ribosyltransferase (polymerizing), ADP-ribosyltransferase 3, ADP-ribosyltransferase C3cer, ADP-ribosyltransferase-1, ADPr, ADPRT, ADPRT , ART, ART1, ARTC2.2, ARTD10, ARTD10/PARP10, ARTD11, ARTD14, ARTD3, ARTD7/PARP15, ARTD8, asparagine-specific ADP-ribosyltransferase, B aggressive lymphoma protein 2, C3 exoenzyme, C3-like toxin, C3larvin toxin, C3lim, CRM66, diphtheria toxin-like ADP-ribosyltranferase, ecto-ADP-ribosyltransferase, ecto-ADP-ribosyltransferase 2.2, ecto-ADP-ribosyltransferase ART2.2, EspJ, Exo53, exoenzyme C3, exoenzyme S, exoenzyme T, ExoS, ExoT, exotoxin-S, mART, ModA, ModB, mono ADP-ribosyltransferase, mono(ADP-ribosyl) transferase, mono-ADP-ribosyl transferase C3, mono-ADP-ribosyltransferase, More, msPARP , NAD(+) ADP-ribosyltransferase , NAD+:ADP-ribosyltransferase (polymerizing), NAD-dependent ADP-ribosyltransferase, NAD-protein ADP-ribosyltransferase, NarE, pADPRT, PAR polymerase, PARP, PARP , PARP-1, PARP-7, PARP-related/IalphaI-related H5/proline-rich , PARP1, PARP10, PARP11, PARP14, PARP3, PARPss, pertussis toxin, PH5P , poly (ADP-ribose) polymerase, poly (ADP-ribose) polymerase-1, poly(ADP-ribose) polymerase, poly(ADP-ribose) polymerase family member 11, poly(ADP-ribose) polymerase-1, poly(ADP-ribose) polymerase-11, poly(ADP-ribose) polymerase-like enzyme, poly(ADP-ribose) synthase, poly(ADP-ribose) transferase, poly(ADP-ribosyl)transferase, poly[ADP-ribose] synthetase , PTx, PtxS1, scabin, TANK1, TANK1 , TANK2 , tankyrase I, Tankyrase-like protein , Tankyrase-related protein , TbSIR2RP1, TiPARP, TNKS-1, TRF1-interacting ankyrin-related ADP-ribose polymerase, TRF1-interacting ankyrin-related ADP-ribose polymerase , TTS-ExoS, type III-secreted toxin, VahC, VPARP , vsdc, YART
ECTree
KM Value
KM Value on EC 2.4.2.30 - NAD+ ADP-ribosyltransferase
Please wait a moment until all data is loaded. This message will disappear when all data is loaded.
Please wait a moment until the data is sorted. This message will disappear when the data is sorted.
0.011 - 0.037
(ADP-D-ribosyl)n-actin
0.017
(ADP-D-ribosyl)n-RhoA protein
at pH 7.4 and 37°C
-
0.03 - 0.429
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor
-
0.0225
N6-etheno-NAD+
-
at pH 7.5 and 22°C
additional information
additional information
-
0.011
(ADP-D-ribosyl)n-actin

mutant N86A, 25°C, pH not specified in the publication
0.02
(ADP-D-ribosyl)n-actin
mutant E213A, 25°C, pH not specified in the publication
0.023
(ADP-D-ribosyl)n-actin
mutant E215A, 25°C, pH not specified in the publication
0.024
(ADP-D-ribosyl)n-actin
wild-type, 25°C, pH not specified in the publication
0.033
(ADP-D-ribosyl)n-actin
mutant S178A, 25°C, pH not specified in the publication
0.035
(ADP-D-ribosyl)n-actin
mutant Y82A, 25°C, pH not specified in the publication
0.037
(ADP-D-ribosyl)n-actin
mutant Y77A, 25°C, pH not specified in the publication
0.03
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor

-
recombinant full-length enzyme
-
0.037
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor
-
recombinant DELTAN222
-
0.049
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor
-
DELTAN222
-
0.23
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor
-
DELTAN222/E381S
-
0.387
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor
-
DELTAN222/E381D
-
0.429
(ADP-D-ribosyl)n-soybean-trypsin-inhibitor
-
DELTAN222/E381A
-
0.002
NAD+

-
-
0.004
NAD+
mutant N86A, 25°C, pH not specified in the publication
0.005
NAD+
mutant E213A, 25°C, pH not specified in the publication
0.006
NAD+
wild-type, 25°C, pH not specified in the publication
0.008
NAD+
mutant Y77A, 25°C, pH not specified in the publication
0.009
NAD+
-
pH 6.0, 30°C, in absence of exosin
0.012
NAD+
mutant Y82A, 25°C, pH not specified in the publication
0.014
NAD+
mutant E215A, 25°C, pH not specified in the publication
0.0171
NAD+
-
pH 7.3, 37°C, cosubstrate RhoC
0.018
NAD+
mutant S178A, 25°C, pH not specified in the publication
0.02
NAD+
-
90000 Da protein
0.02
NAD+
-
hydrolysis of NAD+, wild-type enzyme
0.0208
NAD+
-
pH 7.3, 37°C, cosubstrate RhoA
0.03
NAD+
-
pH 6.0, 30°C, in presence of exosin
0.034
NAD+
at pH 7.4 and 37°C
0.0564
NAD+
-
pH 7.3, 37°C, cosubstrate RhoB
0.068
NAD+
wild type enzyme, at pH 7.9 and 22°C
0.077
NAD+
-
116000 Da protein
additional information
additional information

-
-
-
additional information
additional information
-
kinetics for NAD+ utilization are not Michaelis-Menten type
-




 results (
results ( results (
results ( top
top






