2.7.1.23: NAD+ kinase
This is an abbreviated version!
For detailed information about NAD+ kinase, go to the full flat file.
Reaction
 ATP +
ATP +
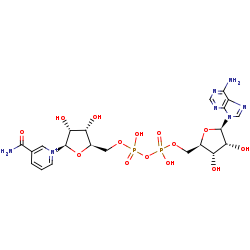 NAD+=
NAD+=
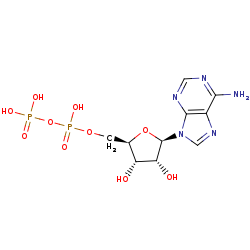 ADP +
ADP +
 NADP+
NADP+
Synonyms
Afnk, AtNADK1, AtNADK2, AtNADK3, ATP: NAD(H) 2'-phosphotransferase, ATP:NAD 2'-phosphotransferase, ATP:NAD+ 2'-phosphotransferase, BsNADK, C5orf33, C5orf33 protein, diphosphopyridine nucleotide kinase, diphosphppyridine kinase, DPN kinase, EC 2.7.1.23, GmNADK1, GmNADK2, GmNADK3, GmNADK4, GmNADK5, HsNADK, kinase (phosphorylating), nicotinamide adenine dinucleotide, kinase, nicotinamide adenine dinucleotide (phosphorylating), MJ0917, NAD kinase, NAD kinase 1, NAD kinase 2, NAD kinase2, NAD(H) kinase, NadF, NADHK, NADK, NADK1, NADK2, NADK3, NADP phosphatase/NAD kinase, native plant calcium- and calmodulin-dependent NAD+-kinase, nicotinamide adenine dinucleotide kinase, OsNADK1, PH1074, Poly(P)/ATP NAD kinase , poly(P)/ATP-NAD kinase, poly(P)/ATPdependent NAD kinase, polyP/ATP-dependent NAD kinase, polyphosphate/ATP-NAD kinase, Pos5, Ppnk, PPNK_THEMA, sll1415, SlNADK1, SlNADK2, slr0040, slr0400, slr1415, SpNADK1, SpNADK2, styNadK, Utr, Utr1, Utr1p, YEF1, Yef1p, YfjB, YjbN, YtdI
ECTree
Turnover Number
Turnover Number on EC 2.7.1.23 - NAD+ kinase
Please wait a moment until all data is loaded. This message will disappear when all data is loaded.
Please wait a moment until the data is sorted. This message will disappear when the data is sorted.
2.3
ADP
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
212
fructose-1,6-bisphosphate
-
1.9
NADH
in 100 mM Tris-HCl (pH 7.5), at 37°C
7.8
polyphosphate(20)
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
7.3
polyphosphate(25)
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
3.4
polyphosphate(45), polyphosphate(65)
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
-
1.8
polyphosphate(75)
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
-
8.7
Tetraphosphate
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
3
Triphosphate
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
additional information
additional information
-
the molecular activity is 360 per min, the activity per active centre is less than 60 per min, pH 7.4, 30°C
-
0.52
ATP

-
wild type enzyme, at pH 7.5 and 30°C
0.6
ATP
Corynebacterium glutamicum subsp. lactofermentum
-
mutant enzyme P57S/P117S, at pH 6.0 and 30°C
2.02
ATP
with NAD+ as cosubstrate, in 100 mM Tris-HCl (pH 7.5), at 37°C
2.54
ATP
Corynebacterium glutamicum subsp. lactofermentum
-
wild type enzyme, at pH 7.5 and 30°C
2.68
ATP
with NADH as cosubstrate, in 100 mM Tris-HCl (pH 7.5), at 37°C
20.51
ATP
Corynebacterium glutamicum subsp. lactofermentum
-
mutant enzyme P57S/P117S, at pH 7.5 and 30°C
29.2
ATP
-
in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
399
ATP
-
85°C, pH 8.5, 20 mM Mg2+
0.41
NAD+

-
mutant D45N, in the presence of 0.5 mM ATP
0.5
NAD+
Corynebacterium glutamicum subsp. lactofermentum
-
mutant enzyme P57S/P117S, at pH 6.0 and 30°C
0.54
NAD+
-
wild type enzyme, at pH 7.5 and 30°C
1.34
NAD+
-
mutant D45N, in the presence of 4 mM ATP
1.9
NAD+
in 100 mM Tris-HCl (pH 7.5), at 37°C
3.62
NAD+
-
wild-type, in the presence of 0.5 mM ATP
3.8
NAD+
Corynebacterium glutamicum subsp. lactofermentum
-
wild type enzyme, at pH 7.5 and 30°C
7.8
NAD+
-
in the presence of polyphosphate(20), in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
13.12
NAD+
-
wild-type, in the presence of 4 mM ATP
15.42
NAD+
Corynebacterium glutamicum subsp. lactofermentum
-
mutant enzyme P57S/P117S, at pH 7.5 and 30°C
27.1
NAD+
-
in the presence of ATP, in 100 mM HEPES, pH 7.3, containing 20 mM MgCl2, at 30°C
424
NAD+
-
85°C, pH 8.5, 20 mM Mg2+


 results (
results ( results (
results ( top
top






