2.3.1.48: histone acetyltransferase
This is an abbreviated version!
For detailed information about histone acetyltransferase, go to the full flat file.
Reaction
 acetyl-CoA +
acetyl-CoA +
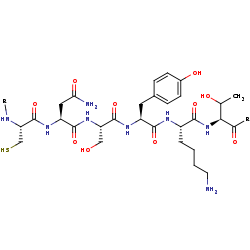 [protein]-L-lysine=
[protein]-L-lysine=
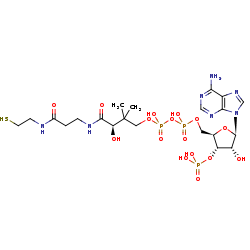 CoA +
CoA +
 [protein]-N6-acetyl-L-lysine
[protein]-N6-acetyl-L-lysine
Synonyms
AAPatA, acetyltransferase, histone, Ada2/Ada3/Gcn5 complex, AIB1, amino acid sensing acetyltransferase, AmiPatA, Amir 5672, ARD1, ARD1 acetyltransferase, arginine methyltransferase A, arginine methyltransferase B, arrest defective 1, arrest defective protein 1 acetyltransferase, At3g14980, ATAC2, ATase1, ATase2, cAMP-regulated protein lysine acetyltransferase, Cbp, CBP histone acetyltransferase, CBP/p300, chameau, circadian locomoter output cycles protein kaput, CLOCK, CREB binding protein, CREB-binding protein, CREBBP, east1, ELO3, ELONGATA3, elongator protein 3, ELP3, enhancer-of-asymmetric leaves-two1, Enok, EP300, Esa1, factor acetyltransferase, FAT, Gcn5, GCN5 family KAT, Gcn5 HAT, GCN5 lysine acetyltransferase, GCN5 N acetyltransferase 5, Gcn5 related N-acetyltransferase, GCN5-like acetyltransferase, GCN5-like enzyme, GCN5-related N-acetyltransferase, Gcn5/PCAF histone acetyltransferase, GCN5b, GCN5L2, general control non-derepressible 5, general control non-repressed protein5, general control nonderepressible 5, general control nonrepressed-protein 5, general control of amino acid synthesis 5, GNAT, GNAT-related histone acetyltransferase complex, GTF3C4, H4 lysine acetyltransferase, HAC1, HAG1, HAG3, HAG4, HAG5, HAM1, HAM2, hARD1, HAT, HAT-B, HAT-B complex, Hat1, Hat1p, Hat2, HatB3.I, HBO1, histone (H4 K16) acetyltransferase, histone acetokinase, histone acetyl transferase, histone acetylase, histone acetyltransferase, histone acetyltransferase 1, histone acetyltransferase AtGCN5, histone acetyltransferase B, histone acetyltransferase Tip60, histone acetyltransferase-1, histone acetyltransferases, histone H3 acetyltransferase, histone H4 acetyltransferase, histone H4 lysine 16 acetyltransferase, histone transacetylase, histone/protein lysine acetyltransferase, HIV-1 Tat-interactive protein, hMOF, Hpa2, Hpa3, human acetylase binding to ORC1, Idm1, K (lysine) acetyltransferase 8, K(lysine) acetyltransferase 8, KAT, KAT10, KAT11, KAT12, KAT13A, KAT13B, KAT13C, KAT13D, KAT2, Kat2A, KAT2A/GCN5, Kat2b, KAT3A, KAT3B, KAT4, KAT5, KAT6, KAT6A, KAT6B, KAT7, KAT8, KAT9, Lsy-12, lysine acetyltransferase, lysine acetyltransferase 2, lysine acetyltransferase 2A, lysine acetyltransferase 2B, lysine acetyltransferase 5, lysine acetyltransferase 8, lysine acetyltransferase complex, lysine acetyltransferase p300, lysine acetyltransferases, lysine-acetyltransferase, Mof, monocytic leukemia zinc finger, monocytic leukemia zinc finger protein, More, Morf, MORF histone acetyltransferase, MOZ, MOZ histone acetyltransferase, Moz related factor, Mst1, MtPat, Myb-binding protein 1A, MYBBP1A, MYST, MYST protein lysine acetyltransferase, MYST-related histone acetyltransferase complex, MYST1, MYST2, MYST3, MYST4, N(alpha)-acetyltransferase 10, N-acetyltransferase, N-terminal acetyltransferase, NAA10, NAT, NAT4, NCoA-1, NCoA-3, NCOAT, Nepsilon-lysine acetyltransferase, NuA4, NuA4 histone acetyltransferase, nuclear cytoplasmicO-GlcNAcase and acetyltransferase, nuclear receptor coactivator 1, nuclear receptor coactivator 3, nucleosome-histone acetyltransferase, Nut1, p/CAF, P/CAF histone acetyltransferase, p160, p300, p300 HAT, p300 histone acetyltransferase, p300/CBP, p300/CBP associated factor, p300/CBP histone acetyltransferase, P300/CBP-associated factor, p300/CBP-associated factor (pCAF), p300HAT, PA4534, Pat, PCAF, PCAF histone acetyltransferase, PF11_0192, PfGCN5, PfGCN5 HAT, Piccolo NuA4 complex, protein acetyl-transferase, protein acetyltransferase, Qkf, regulator of Ty1 transposition protein 109, RmtA, RmtB, RPD3, Rtt109, Rv0998, SAS complex, Sas2, Sas3, SRC1, SRC3, Sven 0867, SvePatA, TAF, TAF1, Tat interacting protein of 60 kDa, Tat-interactive protein, 60 kDa, TBP1, TFIIIC90, tGCN5, TgGCN5b, TgMYST-B, Tip60, tumor suppressor histone H3 lysine 23 acetyltransferase, type B histone acetyltransferase, YfmK, zMoz
ECTree
KM Value
KM Value on EC 2.3.1.48 - histone acetyltransferase
Please wait a moment until all data is loaded. This message will disappear when all data is loaded.
Please wait a moment until the data is sorted. This message will disappear when the data is sorted.
0.0002 - 0.0086
3-azidopropionyl-CoA
0.0009 - 0.0024
4-pentynoyl-CoA
0.0006
5-hexynoyl-CoA
mutant GCN5 T612G, pH and temperature not specified in the publication
0.0003
6-heptynoyl-CoA
mutant GCN5 T612G, pH and temperature not specified in the publication
0.0002 - 0.046
acetyl-CoA
0.007 - 2.09
histone H3
-
0.044 - 0.112
histone H3 tail peptide
-
0.05 - 0.49
histone H3-peptide
-
1.12
histone H3-peptide p19
-
PCAF catalytic domain
-
0.75
histone H3-peptide p27
-
PCAF catalytic domain
-
0.0000053 - 0.5
histone H4
0.0208 - 0.197
histone H4 peptide
-
0.135 - 0.372
piccoloNuA4 peptide
-
1.28 - 4.63
protein p53
-
0.0081
SGRGKGGKGLGKGGAKRHRK
at pH 8.0 and 30°C
additional information
additional information
-
0.0002
3-azidopropionyl-CoA

mutant GCN5 T612G, pH and temperature not specified in the publication
0.0082
3-azidopropionyl-CoA
wild-type GCN5, pH and temperature not specified in the publication
0.0086
3-azidopropionyl-CoA
pH and temperature not specified in the publication
0.0009
4-pentynoyl-CoA

mutant GCN5 T612G, pH and temperature not specified in the publication
0.0024
4-pentynoyl-CoA
pH and temperature not specified in the publication
0.0002
acetyl-CoA

-
enzyme form B
0.00023
acetyl-CoA
-
enzyme form DB
0.00027
acetyl-CoA
-
pH 7.5, 25°C, mutants D639E and L503P/D601G/Y612C, with substrate histone H3
0.00028
acetyl-CoA
-
pH 7.5, 25°C, wild-type P/CAF enzymem, with substrate histone H3
0.00029
acetyl-CoA
-
pH 7.5, 25°C, mutant Y612C/P655R, with substrate histone H3
0.0003
acetyl-CoA
wild-type, pH 7.5, 25°C
0.00033
acetyl-CoA
-
pH 7.5, 25°C, mutant L503P, with substrate histone H3
0.00034
acetyl-CoA
-
pH 7.5, 25°C, mutants D601G and V582A/D639E, with substrate histone H3
0.00036
acetyl-CoA
-
pH 7.5, 25°C, mutant V582A, with substrate histone H3
0.00041
acetyl-CoA
-
pH 7.5, 25°C, mutant Y612C, with substrate histone H3
0.00049
acetyl-CoA
mutant D288N, pH 7.5, 25°C
0.00054
acetyl-CoA
at pH 8.0 and 30°C
0.00058
acetyl-CoA
-
Gcn5
0.00062
acetyl-CoA
-
Gcn5
0.0007
acetyl-CoA
-
Gcn5 mutant A190T
0.0007
acetyl-CoA
mutant D287N, pH 7.5, 25°C
0.00098
acetyl-CoA
-
PCAF protein
0.0011
acetyl-CoA
at 0.015 mM histone H4 peptide, pH and temperature not specified in the publication
0.0012
acetyl-CoA
pH 7.5, 25°C, mutant E338Q
0.0012
acetyl-CoA
at 0.030 mM histone H4 peptide, pH and temperature not specified in the publication
0.0015
acetyl-CoA
pH 7.5, 25°C, mutant C304A
0.0016
acetyl-CoA
-
recombinant PCAF catalytic domain mutant Y638A
0.0016
acetyl-CoA
-
recombinant wild-type PCAF catalytic domain
0.002
acetyl-CoA
pH 7.5, 25°C, mutant C304S
0.0021
acetyl-CoA
-
Gcn5 mutant A190S
0.0021
acetyl-CoA
at 0.060 mM histone H4 peptide, pH and temperature not specified in the publication
0.0023
acetyl-CoA
-
wild-type, presence of 0.4 equivalents of chaperone Vps75, pH 7.5, 25°C
0.0025
acetyl-CoA
-
Gcn5 protein
0.0025
acetyl-CoA
pH 7.5, 25°C, wild-type enzyme
0.0028
acetyl-CoA
at 0.090 mM histone H4 peptide, pH and temperature not specified in the publication
0.0033
acetyl-CoA
-
autoacetylation
0.0037
acetyl-CoA
wild-type GCN5, pH and temperature not specified in the publication
0.0039
acetyl-CoA
-
wild-type, presence of 0.2 equivalents of chaperone Vps75, pH 7.5, 25°C
0.004
acetyl-CoA
mutant GCN5 T612G, pH and temperature not specified in the publication
0.0063
acetyl-CoA
-
wild-type, presence of 1 equivalent of chaperone Vps75, pH 7.5, 25°C
0.0065
acetyl-CoA
-
wild-type, presence of 2 equivalents of chaperone Vps75, pH 7.5, 25°C
0.0065
acetyl-CoA
-
wild-type, presence of 8 equivalents of chaperone Vps75, pH 7.5, 25°C
0.0067
acetyl-CoA
wild-type, pH not specified in the publication, temperature not specified in the publication
0.01
acetyl-CoA
-
+ spermidine, enzyme form B
0.015
acetyl-CoA
-
+ spermidine, enzyme form A
0.0173
acetyl-CoA
pH and temperature not specified in the publication
0.046
acetyl-CoA
-
PCAF catalytic domain
0.007
histone H3

wild-type, pH 7.5, 25°C
-
0.039
histone H3
-
pH 7.5, 25°C, mutant D639E
-
0.053
histone H3
-
pH 7.5, 25°C, wild-type P/CAF enzyme
-
0.054
histone H3
-
pH 7.5, 25°C, mutant L503P
-
0.055
histone H3
-
pH 7.5, 25°C, mutant D601G
-
0.062
histone H3
-
pH 7.5, 25°C, mutant Y612C/P655R
-
0.064
histone H3
-
pH 7.5, 25°C, mutant L503P/D601G/Y612C
-
0.165
histone H3
-
pH 7.5, 25°C, mutant Y612C
-
0.24
histone H3
wild-type, pH 8.0, temperature not specified in the publication
-
0.273
histone H3
-
Gcn5 mutant A190S
-
0.352
histone H3
-
Gcn5 mutant A190T
-
0.357
histone H3
-
Gcn5
-
0.43
histone H3
mutant having m-fluorophenylalanine incorporated, pH 8.0, temperature not specified in the publication
-
0.46
histone H3
-
pH 7.5, 25°C, mutant V582A/D639E
-
0.471
histone H3
-
Gcn5
-
0.532
histone H3
-
PCAF
-
0.56
histone H3
-
pH 7.5, 25°C, mutant V582A
-
0.69
histone H3
-
recombinant p-fluorophenylalanine-substituted tGCN5
-
0.7
histone H3
mutant having p-fluorophenylalanine incorporated, pH 8.0, temperature not specified in the publication
-
0.8
histone H3
-
recombinant o-fluorophenylalanine-substituted wild-type tGCN5
-
1.05
histone H3
-
recombinant wild-type tGCN5
-
2.09
histone H3
-
recombinant m-fluorophenylalanine-substituted wild-type tGCN5
-
0.044
histone H3 tail peptide

mutant D287A/D288A, pH 7.5, 25°C
-
0.055
histone H3 tail peptide
mutant D288N, pH 7.5, 25°C
-
0.062
histone H3 tail peptide
mutant D287N, pH 7.5, 25°C
-
0.112
histone H3 tail peptide
wild-type, pH 7.5, 25°C
-
0.05
histone H3-peptide

-
PCAF catalytic domain
-
0.49
histone H3-peptide
-
Gcn5 protein
-
0.0000053
histone H4

-
in the absence of heparin
0.000259
histone H4
-
in the presence of heparin
0.11
histone H4
-
pH 7.5, 25°C, mutant L503P/D601G/Y612C
0.12
histone H4
-
pH 7.5, 25°C, mutant Y612C
0.14
histone H4
-
pH 7.5, 25°C, mutant Y612C/P655R
0.15
histone H4
-
pH 7.5, 25°C, mutant D639E
0.19
histone H4
-
pH 7.5, 25°C, wild-type P/CAF enzyme
0.21
histone H4
-
pH 7.5, 25°C, mutant L503P
0.25
histone H4
-
pH 7.5, 25°C, mutant D601G
0.31
histone H4
-
pH 7.5, 25°C, mutant V582A/D639E
0.5
histone H4
-
pH 7.5, 25°C, mutant V582A
0.0208
histone H4 peptide

wild-type, pH not specified in the publication, temperature not specified in the publication
-
0.0595
histone H4 peptide
mutant W199A, pH not specified in the publication, temperature not specified in the publication
-
0.086
histone H4 peptide
mutant D62A, pH not specified in the publication, temperature not specified in the publication
-
0.099
histone H4 peptide
mutant D277N, pH not specified in the publication, temperature not specified in the publication
-
0.1
histone H4 peptide
mutant E276Q, pH not specified in the publication, temperature not specified in the publication
-
0.1267
histone H4 peptide
mutant E64A, pH not specified in the publication, temperature not specified in the publication
-
0.197
histone H4 peptide
mutant E187Q, pH not specified in the publication, temperature not specified in the publication
-
0.135
piccoloNuA4 peptide

pH 7.5, 25°C, mutant E338Q
-
0.182
piccoloNuA4 peptide
pH 7.5, 25°C, mutant C304S
-
0.216
piccoloNuA4 peptide
pH 7.5, 25°C, wild-type enzyme
-
0.372
piccoloNuA4 peptide
pH 7.5, 25°C, mutant C304A
-
1.28
protein p53

mutant having p-fluorophenylalanine incorporated, pH 8.0, temperature not specified in the publication
-
4.63
protein p53
wild-type, pH 8.0, temperature not specified in the publication
-
0.18
spermidine

-
enzyme form B
0.27
spermidine
-
enzyme form A
additional information
additional information

-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
-
-
additional information
additional information
-
Gcn5 protein mutants
-
additional information
additional information
Michaelis-Menten kinetics
-
additional information
additional information
Michaelis-Menten kinetics
-
additional information
additional information
-
kinetics, site-specific kinetic analysis of loop autoacetylation, PCAF-mediated autoacetylation, overview
-
additional information
additional information
steady-state kinetic analyses, kinetic mechanism
-
additional information
additional information
kinetics and catalytic mechanism of the bi-substrate enzyme KAT8, non-linear Michaelis-Menten regression, overview
-
additional information
additional information
kinetics and catalytic mechanism of the bi-substrate enzyme KAT8, non-linear Michaelis-Menten regression, overview
-




 results (
results ( results (
results ( top
top






